Revealing the Fluctuations of Flexible DNA in 3D
First-of-their-kind images could aid in use of DNA to build tiny, lightweight devices.
The Science
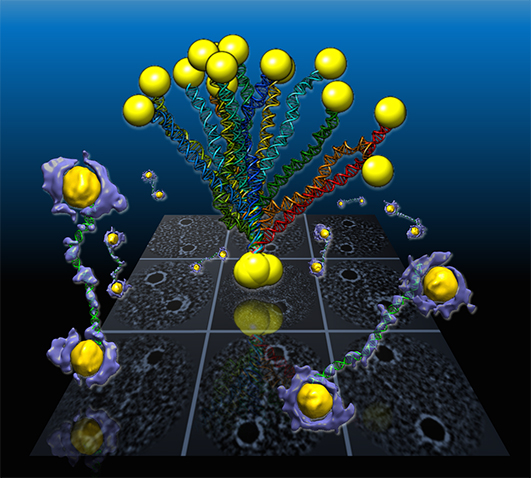
For the first time, an international team of scientists captured high-resolution 3D images from individual double-helix DNA segments attached at either end to gold nanoparticles. The images detail the flexible structure of the DNA segments, which appear as tiny jump ropes.
The Impact
This unique imaging capability should lead to better understanding of disease-relevant proteins and the assembly process that forms DNA. It could also aid in the use of DNA segments as biomarkers, drug-delivery systems, computer memory, and electronic devices.
Summary
Working at the Molecular Foundry using a cutting-edge electron microscope technique coupled with a protein-staining process and sophisticated software, scientists reconstructed the shapes of the coiled DNA strands, which were sandwiched between polygon-shaped gold nanoparticles. The flexible structure of the DNA segments appear as nanoscale jump ropes. The scientists examined the strands with a unique imaging capability they developed at the Molecular Foundry. The technique is called individual-particle electron tomography (IPET). It offers structural details at the scale of about 2 nanometers, or two billionths of a meter. The technique could aid in the use of DNA segments as building blocks in nanoscale drug-delivery systems, markers for biological research, and components for computer memory and electronic devices. It could also lead to images of important disease-relevant proteins that have proven elusive for other imaging techniques and of the assembly process that forms DNA from separate, individual strands.
Contact
Gang (Gary) Ren
Molecular Foundry, Berkeley Lab
gren@lbl.gov; 510.495.2375
Funding
This material is based upon work supported by the National Science Foundation under grant DMR-1344290. Work at the Molecular Foundry was supported by the Office of Science, Office of Basic Energy Sciences of the U.S. Department of Energy under contract no. DE-AC02-05CH11231. G.R. is supported by the National Heart, Lung, and Blood Institute of the National Institutes of Health (no. R01HL115153) and the National Institute of General Medical Sciences of the National Institutes of Health (no. R01GM104427). L.Z. is partly supported by National Natural Science Foundation of China (no. 11504287) and Young Talent Support Plan of Xi’an Jiaotong University; Z.L. and J.L. are partly supported by the National Basic Research Program of Ministry of Science and Technology, China (no. 2015CB553602).
Publications
L. Zhang, D. Lei, J. M. Smith, M. Zhang, H. Tong, X. Zhang, Z. Lu, J. Liu, A. P. Alivisatos, and G. Ren. “Three-dimensional structural dynamics and fluctuations of DNA-nanogold conjugates by individual-particle electron tomography.” Nature Communications 7:11083 (2016). [DOI: 10.1038/ncomms11083]
Related Links
LBL | Revealing the Fluctuations of Flexible DNA
Highlight Categories
Performer: DOE Laboratory , SC User Facilities , Foundry , BES User Facilities
Additional: Collaborations , Non-DOE Interagency Collaboration , International Collaboration



